OpenDRC: An Open-Source Implementation |
DRCFilter allows the researcher to extract the activation information from one or more ACTS file and will output this information in a form suitable for inclusion in MS Excel or Matlab so that it can be graphed. Separate Matlab routines are also included in OpenDRC1.2 that will display the activations in a graph for the Matlab user. These can be used as a starting point for researchers own Matlab programs.
DRCFilter Examples:
- DRCFilter used to filter all the LL and GPC activations from one input ACTS file (.acts) to the output text file (.txt):
... and the results look like this (NOTE: The right-hand-side of the output has been trimmed to allow it to fit this page):
DRCFilter -f<test-word.acts> -wL*:* -wGPC*:* -ot<test-word.txt>
11 73 14 ; DRCFilter v1.0.0 ; Run date, time: Thu 26/May/2011 23:33:42 ; NOTE: Numbers on first line are: Header-Lines, Data-Lines, and Activation-columns ; Command line arguments: ; -fbelch-1.drc\belch.acts ; -wL*:* ; -wGPC*:* ; -otBelch.txt ; Cycle Range: [1,73] ; Cycle Limits: [-1,-1] Cycle F01-LL0:b F01-LL1:e F01-LL2:l F01-LL3:c F01-LL4:h F01-LL5:+ ... 1 0.014000 0.014000 0.014000 0.014000 0.014000 0.014000 ... 2 0.027804 0.027804 0.027804 0.027804 0.027804 0.027804 3 0.041415 0.041415 0.041415 0.041415 0.041415 0.041415 4 0.054835 0.054835 0.054835 0.054835 0.054835 0.054835 5 0.068067 0.068067 0.068067 0.068067 0.068067 0.068067 6 0.081114 0.081114 0.081114 0.081114 0.081114 0.081114 7 0.093979 0.093979 0.093979 0.093979 0.093979 0.093979 ... 8 0.106729 0.106729 0.106729 0.106729 0.106729 0.106729 9 0.119442 0.119442 0.119442 0.119442 0.119442 0.119442 10 0.132188 0.132188 0.132188 0.132188 0.132188 0.132188 11 0.145034 0.145034 0.145034 0.145034 0.145034 0.145034 12 0.158041 0.158041 0.158041 0.158041 0.158041 0.158041 ... 13 0.171264 0.171264 0.171264 0.171264 0.171264 0.171264 14 0.184755 0.184755 0.184755 0.184755 0.184755 0.184755 15 0.198558 0.198558 0.198558 0.198558 0.198558 0.198558 16 0.212713 0.212713 0.212713 0.212713 0.212713 0.212713 17 0.227256 0.227256 0.227256 0.227256 0.227256 0.227256 18 0.242214 0.242214 0.242214 0.242214 0.242214 0.242214 ... ...The first line of the output file is
lines-of-header lines-of-data number-of-data-fields. This line is used by the given Matlab program to strip the data out of the file and to properly label the activation lines in the legend. The lines starting with a semicolon are comment lines for reference. The line starting with "Cycle" contains the column headers (explained below) and the remaining lines (starting with numbers) are the data lines.When the above output is run through the OpenDRC1.2 Matlab example graphing routines the output is as follows:
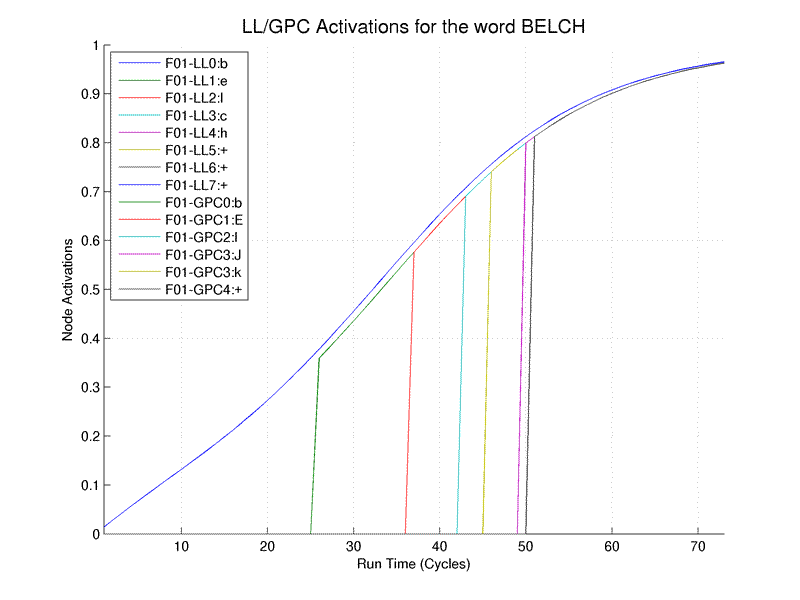
The legend and the data-field headers require some explanation. The first part (e.g. F01-) indicates the input file number from which the data was extracted. This is important as activation curves can be extracted from multiple files.
The second part (e.g. LL0:b) is <layer><column or node>:<word, letter or phoneme>. Thus F01-LL0:b indicates the activation curve was taken from the first file and the 0th column of the Letter Layer and the active letter is 'b'.
- To run DRCFilter from more than one file we just have to specify the input file and the 'watches'. In the following example the third (numbering starts at zero) column letters from the Letter-Layer and phonemes from the GPC-Layer are extracted from the first file and the fourth from the second file.
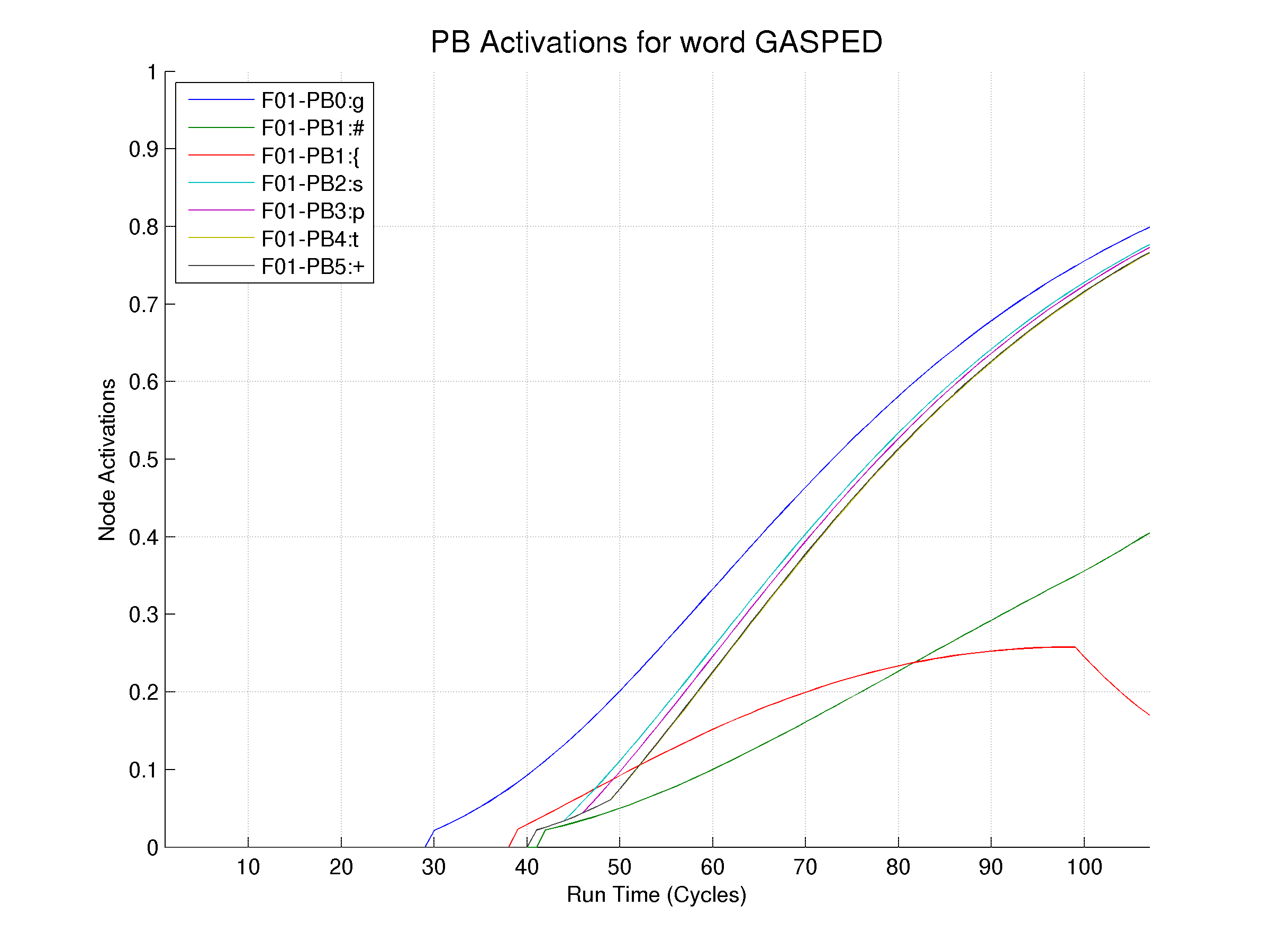
DRCFilter -f<test-word1.acts> -wL2:* -wGPC2:* -f<test-word2.acts> -wL3:* -wGPC3:* -ot<test-word.txt>- The following is an interesting example from the word GASPED watching the Phoneme Buffer activations. GASPED has a WAMMIE (see Rastle and Coltheart, 1998) in the second column of the Phoneme Buffer (PB1). The original phoneme F01-PB1:#(red) is WAMMIED at approximately cycle 100 and F01-PB1:{ takes over(green)
DRCFilter -fgasped.drc\gasped.acts -wP*:* -otgasped.txtThis produces the following output file:
10 107 7 ; DRCFilter v1.0.0 ; Run date, time: Thu 07/Apr/2011 15:17:23 ; NOTE: Numbers on first line are: Header-Lines, Data-Lines, and Activation-columns ; Command line arguments: ; -fgasped.drc\gasped.acts ; -wP*:* ; -otgasped.txt ; Cycle Range: [1,107] ; Cycle Limits: [-1,-1] Cycle F01-PB0:g F01-PB1:# F01-PB1:{ F01-PB2:s F01-PB3:p F01-PB4:t F01-PB5:+ 1 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 2 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 3 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 4 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 5 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 ... 95 0.719434 0.324671 0.257038 0.687747 0.682910 0.672958 0.674077 96 0.727036 0.331020 0.257480 0.696208 0.691502 0.681814 0.682905 97 0.734452 0.337344 0.257786 0.704461 0.699883 0.690451 0.691516 98 0.741683 0.343642 0.257957 0.712509 0.708056 0.698875 0.699914 99 0.748733 0.349915 0.257993 0.720355 0.716024 0.707088 0.708101 100 0.755606 0.356162 0.245061 0.728004 0.723790 0.715095 0.716083 101 0.762303 0.362627 0.232736 0.735458 0.731360 0.722898 0.723861 102 0.768829 0.369288 0.220988 0.742721 0.738736 0.730503 0.731441 103 0.775188 0.376127 0.209794 0.749798 0.745922 0.737912 0.738826 104 0.781381 0.383124 0.199126 0.756691 0.752922 0.745129 0.746020 105 0.787414 0.390261 0.188963 0.763405 0.759740 0.752159 0.753027 106 0.793289 0.397521 0.179281 0.769943 0.766379 0.759006 0.759850 107 0.799009 0.404888 0.170060 0.776309 0.772844The above file run through the Matlab routines produces the following plot:
DRCFilter help:
OpenDRC homeHelp for DRCFilter can be obtained by issuing the command
DRCFilter -hand will display:DRCFilter v1.0.0 Copyright (C) 2011 Alan Angold This program comes with ABSOLUTELY NO WARRANTY. This is free software, and you are welcome to redistribute it under certain conditions. See the included file 'GNU General Public License, version 3 _GPLv3_.pdf' for details. OpenDRC URL: https://sourceforge.net/projects/opendrc/ Usage: DRCFilter {-P<pfile>|-f<afile>|-c<min>:<Max>|-w<LID>[:<NID>]|-o[t|x]<ofile>|-h|-d} -P Open file <pfile> from which to read command line arguments File can include any -P,-f,-w,-c argumens, one per line -f Open ACTS file <afile> to read following activations from -c Output ACTS cycles <min> to <Max> -o Output the results to the Text(t) or EXCEL(X) file <ofile> -h This help message -d Dump Watches (debug) -w Watch a particular activation (LID=Layer ID,NID=Node ID, cid ranges 1-N, '*'-match all): LID NID VF<cid> - Features (0-13) in column cid to watch L<cid> - Letter (a-z,+) in column cid to watch Orth - Word in OIL to watch Phon - Phonemic word in POL to watch GPC<cid> - Phoneme in GPC column cid to watch GPCR<cid> - Phoneme in GPCR column cid to watch P<cid> - Phoneme in PB column cid to watch TL TO TP TGPC TGPCR TPh Examples: To watch the letter 'b' in the third column of the letter level: "-wL3:b". To watch all the phonemes in the PB in column 1: "-wP1:*". To watch all the (active) words in the OIL: "-wOrth:*". To watch the POL totals: "-wTP". Examples: To track the activations of all the phonemic words in the POL for the word "beech" use the command: "DRCFilter -fbeech.acts -wPhon:*" To watch all letter activations in all the colmns of the Letter Level(LL) from two runs of OpenDRC/DRC1.2 for the word "jump": "DRCFilter -f\jump.drc\jump.acts -wL*:* -f\jump-1.drc\jump.acts -wL*:*